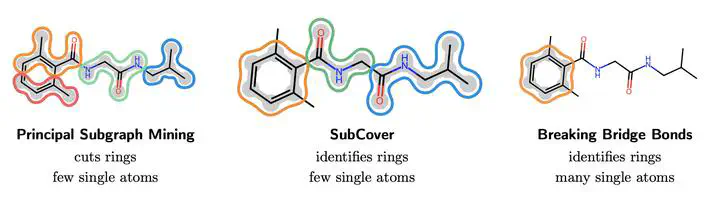
 Concept Figure
Concept Figure
Abstract
Machine learning for molecules holds great potential for efficiently exploring the vast chemical space and thus streamlining the drug discovery process by facilitating the design of new therapeutic molecules. Deep generative models have shown promising results for molecule generation, but the benefits of specific inductive biases for learning distributions over small graphs are unclear. Our study aims to investigate the impact of subgraph structures and vocabulary design on distribution learning, using small drug molecules as a case study. To this end, we introduce Subcover, a new subgraph-based fragmentation scheme, and evaluate it through a two-step variational auto-encoder. Our results show that Subcover’s improved identification of chemically meaningful subgraphs leads to a relative improvement of the FCD score by 30%, outperforming previous methods. Our findings highlight the potential of Subcover to enhance the performance and scalability of existing methods, contributing to the advancement of drug discovery.