 Graphs in single-cell genomics
Graphs in single-cell genomics
Abstract
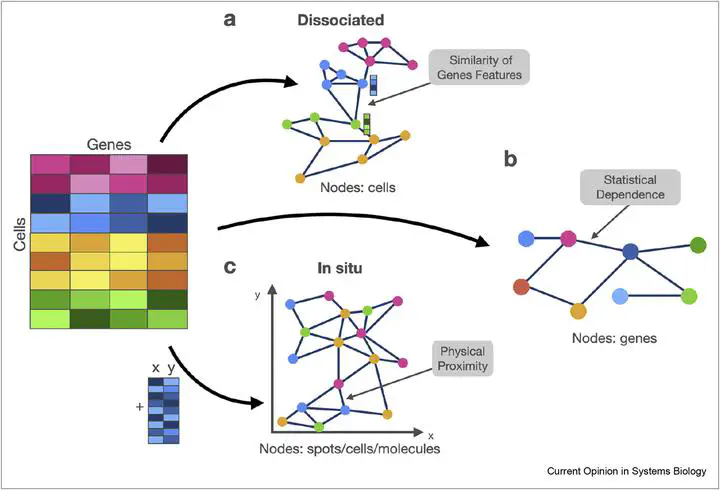
Single-cell RNA sequencing measures gene expression at an unprecedented resolution and scale and allows the analysis of cellular phenotypes which was not possible before. In this context, graphs occur as a natural representation of the system —both as gene-centric and cell-centric. However, many advances in machine learning on graphs are not yet harnessed in models on single-cell data. Taking the inference of cell types or gene interactions as examples, graph representation learning has a wide applicability to both cell and gene graphs. Recent advances in spatial molecular profiling additionally put graph learning in the focus of attention because of the innate resemblance of spatial information to spatial graphs. We argue that graph embedding techniques have great potential for various applications across single-cell biology. Here, we discuss how graph representation learning maps to current models and concepts used in single-cell biology and formalise overlaps to developments in graph-based deep learning.